Note
Go to the end to download the full example code.
Gaussian Bures-Wasserstein barycenters
Note
Example added in release: 0.9.2.
Illustration of Gaussian Bures-Wasserstein barycenters.
# Authors: Rémi Flamary <remi.flamary@polytechnique.edu>
#
# License: MIT License
# sphinx_gallery_thumbnail_number = 2
Define Gaussian Covariances and distributions
Plot the distributions
def draw_cov(mu, C, color=None, label=None, nstd=1):
def eigsorted(cov):
vals, vecs = np.linalg.eigh(cov)
order = vals.argsort()[::-1]
return vals[order], vecs[:, order]
vals, vecs = eigsorted(C)
theta = np.degrees(np.arctan2(*vecs[:, 0][::-1]))
w, h = 2 * nstd * np.sqrt(vals)
ell = Ellipse(
xy=(mu[0], mu[1]),
width=w,
height=h,
alpha=0.5,
angle=theta,
facecolor=color,
edgecolor=color,
label=label,
fill=True,
)
pl.gca().add_artist(ell)
# pl.scatter(mu[0],mu[1],color=color, marker='x')
axis = [-1.5, 5.5, -1.5, 5.5]
pl.figure(1, (8, 2))
pl.clf()
pl.subplot(1, 4, 1)
draw_cov(m1, C1, color="C0")
pl.axis(axis)
pl.title("$\mathcal{N}(m_1,\Sigma_1)$")
pl.subplot(1, 4, 2)
draw_cov(m2, C2, color="C1")
pl.axis(axis)
pl.title("$\mathcal{N}(m_2,\Sigma_2)$")
pl.subplot(1, 4, 3)
draw_cov(m3, C3, color="C2")
pl.axis(axis)
pl.title("$\mathcal{N}(m_3,\Sigma_3)$")
pl.subplot(1, 4, 4)
draw_cov(m4, C4, color="C3")
pl.axis(axis)
pl.title("$\mathcal{N}(m_4,\Sigma_4)$")
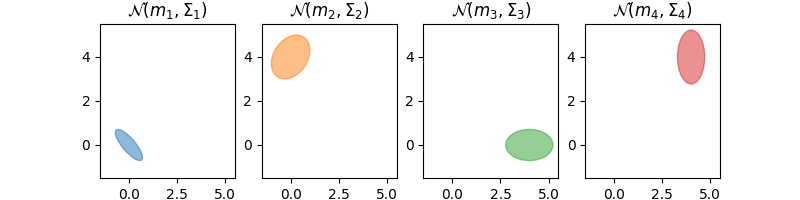
Text(0.5, 1.0, '$\\mathcal{N}(m_4,\\Sigma_4)$')
Compute Bures-Wasserstein barycenters and plot them
# basis for bilinear interpolation
v1 = np.array((1, 0, 0, 0))
v2 = np.array((0, 1, 0, 0))
v3 = np.array((0, 0, 1, 0))
v4 = np.array((0, 0, 0, 1))
colors = np.stack(
(colors.to_rgb("C0"), colors.to_rgb("C1"), colors.to_rgb("C2"), colors.to_rgb("C3"))
)
pl.figure(2, (8, 8))
nb_interp = 6
for i in range(nb_interp):
for j in range(nb_interp):
tx = float(i) / (nb_interp - 1)
ty = float(j) / (nb_interp - 1)
# weights are constructed by bilinear interpolation
tmp1 = (1 - tx) * v1 + tx * v2
tmp2 = (1 - tx) * v3 + tx * v4
weights = (1 - ty) * tmp1 + ty * tmp2
color = np.dot(colors.T, weights)
mb, Cb = ot.gaussian.bures_wasserstein_barycenter(m, C, weights)
draw_cov(mb, Cb, color=color, label=None, nstd=0.3)
pl.axis(axis)
pl.axis("off")
pl.tight_layout()

Total running time of the script: (0 minutes 0.348 seconds)